Process class definition: A process is defined through the attributes name, flux, source, target, delta, sp_at_formation, atomic_fraction, epsilon and epsilon_sp. More...
Public Member Functions | |
| __init__ (self, _process, _data) | |
| progress (self) | |
| Called after each handling step in order to keep track of data access. | |
| set_timeresolution (self, _timeresolution) | |
| Scale data depending on available and required time resolution. | |
| initialise (self, _timeresolution) | |
| Converts Sets time resolution. | |
| deposition (self, _flux, _target, _dt, _f) | |
| Deposition method, used for isotope mixing: This method is specifically designed for a situation in which a nutrient exclusively enters the model domain and mixes with the target pool. | |
| mixingpools (self, _flux, _source, _ds, _target, _dt) | |
| Mixingpools method: This method is a more general implemetation of isotpe mixing, but eventually always calls the method deposition. | |
| reactionpools (self, _flux, _source, _ds, _target, _dt) | |
| Reactionpools method, calculates isotope fractionation: | |
| handle (self, _pools) | |
| Handle method, triggers appropriate isotope calculation: | |
| convert_delta_to_atomic_fraction (self) | |
| get_dimensions (self, _pools, _entities="both", _offset=0) | |
Detailed Description
Process class definition: A process is defined through the attributes name, flux, source, target, delta, sp_at_formation, atomic_fraction, epsilon and epsilon_sp.
name is used to read initial mass from the initialisation file specified in config.json
flux is the amount of nutrient converted from source to target
source and target give the pools from (to) which the amount of nutrient given in flux is transformed or mixed
delta The 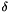
sp_at_formation exclusively applies for processes that produce 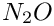
atomic_fraction Atom fraction
epsilon represents the (net) isotpe effect for a given process
epsilon_sp represents the (net) isotpe effect on SP for reduction of 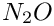
Member Function Documentation
◆ deposition()
| pysimone.process_t.deposition | ( | self, | |
| _flux, | |||
| _target, | |||
| _dt, | |||
| _f ) |
Deposition method, used for isotope mixing: This method is specifically designed for a situation in which a nutrient exclusively enters the model domain and mixes with the target pool.
Since for this special case, the direction of the flux is unequivocal, and the source is outside the model domain, a case distinction as for the mixingpools method is not necessary.
_flux is the amount of compound mixed
_target is the target pool name
_dt is the soil layer?? why is it sometimes _d and then _dt
_f is the atom fraction of the flux, i.e., the atom fraction of the source pool
◆ handle()
| pysimone.process_t.handle | ( | self, | |
| _pools ) |
Handle method, triggers appropriate isotope calculation:
This method eventually decides if mixing or fractiontion is calculated. In addition, it considers that the parent biogeochemical model distributes nutrients via the management option tilling as well as leaching and diffusion. This redistribtion / transport is also covered by the handle method and applies either to the whole profile or to the part of the soil profile affected by tilling. _flux flux for the specific process and the specific time step _pools Dictionary of pools defined in model.json
◆ mixingpools()
| pysimone.process_t.mixingpools | ( | self, | |
| _flux, | |||
| _source, | |||
| _ds, | |||
| _target, | |||
| _dt ) |
Mixingpools method: This method is a more general implemetation of isotpe mixing, but eventually always calls the method deposition.
A more general implementation is required because the direction of a flux between two pools is a priori not clear. The direction of the flux depends on its sign and, thus, determines which pool is the source and which one is the traget. This applies specifically to diffusion or transport of nutrients between aerobic an anaerobic soil compartments. In addition, this method also limits fluxes out of pools to the actual pool size. It is important to note that in such a situation, the isotopic composition is set to 0, which is not accurate, but does not affect later mixing processes as the pool mass is 0.
Please note that mixingpools is always called with an assumption what source and target are, but based on the case selection, the deposition function is eventually called with either the _source OR _target parameter of the mixingpools function.
_flux represents the amount of substrate (source) compound transformed into product (target)
_source represents the name of the source pool
_ds represents the layer of the source
_target represents the name of the target pool
_dt represents the layer of the target pool
◆ reactionpools()
| pysimone.process_t.reactionpools | ( | self, | |
| _flux, | |||
| _source, | |||
| _ds, | |||
| _target, | |||
| _dt ) |
Reactionpools method, calculates isotope fractionation:
This method calculates isotopic fractionation but also has to consider quantitative pool transformation. Quantitative pool transformation results in mixing of the existing source pool to the target pool (this is the first scenario for a positive flux). The second scenario contains the actual calculation of isotopic fractionation based on Rayleigh distillation. It is important to note that this method requires a layer for source and target because some fractionation processes occurr across layer boundaries. Specifically, plants are represented by one single layer, but are capable of taking up nutrients from any layer, with uptake discriminating against the heavy isotope.
_flux represents the amount of substrate (source) compound transformed into product (target)
_source represents the name of the source pool
_ds represents the layer of the source
_target represents the name of the target pool
_dt represents the layer of the target pool
The documentation for this class was generated from the following file:
Generated by