 |
LandscapeDNDC 1.37.0
|
 |
LandscapeDNDC 1.37.0
|
Vegetation model PNET. More...
#include <models/physiology/pnet/pnet.h>
Public Member Functions | |
| lerr_t | solve () |
| lerr_t | sleep () |
Private Member Functions | |
| void | update_nitrogen_concentration (MoBiLE_Plant *) |
| void | YearlyReset (MoBiLE_Plant *) |
| void | Phenology (MoBiLE_Plant *, int, double) |
| void | Photosyn (MoBiLE_Plant *) |
| void | AllocateMo (MoBiLE_Plant *) |
| void | AllocateYr (MoBiLE_Plant *) |
| void | N_uptake (MoBiLE_Plant *, double &) |
| void | Update (MoBiLE_Plant *, double, double) |
| void | TransformPhysiology (MoBiLE_Plant *) |
| size_t | get_na_max_pnet () |
Static Private Attributes | |
| static const double | TMIN_NUP = 6.0 |
| static const double | TMAX_NUP = 25.0 |
| static const double | KM_NO3 = 1.0 |
| static const double | KM_NH4 = 1.0 |
| static const double | SRATEMAX = 0.01 |
| static const double | C = 0.02 |
| static const double | NUPTAIR = 0.0 |
| static const double | GROWMAX = 0.95 |
Vegetation model PNET.
|
private |
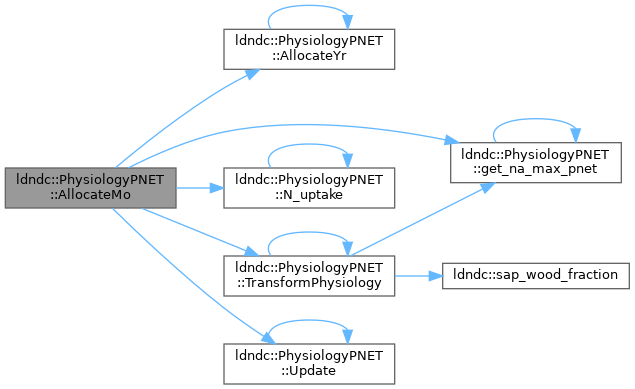
AllocateMo
References AllocateYr(), get_na_max_pnet(), N_uptake(), TransformPhysiology(), and Update().
Referenced by solve().

|
private |
AllocateYr
NOTE: the shift in carbon between compartments disturbs the nitrogen balance at this day!!!
References AllocateYr(), and GROWMAX.
Referenced by AllocateMo(), AllocateYr(), and solve().


|
private |
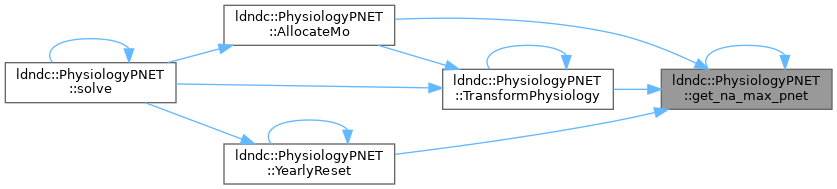
get_na_max_pnet
References get_na_max_pnet().
Referenced by AllocateMo(), get_na_max_pnet(), TransformPhysiology(), and YearlyReset().
|
private |
N_uptake
References KM_NH4, KM_NO3, N_uptake(), TMAX_NUP, and TMIN_NUP.
Referenced by AllocateMo(), N_uptake(), and solve().


|
private |
Phenology
References Phenology().
Referenced by Phenology(), and solve().

|
private |
Photosyn
NOTE: This module is written for photosynthesis responses to monthly average climate values and should be used with care if applied on an daily time step. In PNET-N-DNDC the available light is reduced by: PAR_f = par / (Lai * 0.1 + 1.0); and the light response is changed to: LightEff = (1.0 - pow(2.666, (-Il / HALFSAT[p]))); If the Lai-impact on LightEff had not been limited to Lai 15, the responses would be equal at Lai app. 22. As it is, the photosynthesis in PNET is always larger (app. twice as high).
CHANGES:
AMAXA and AMAXB are specific for the PnET module. The parameters define the photosynthesis rate in dependence on nitrogen, assuming a linear relationship. AMAXA is a mere hypothetical photosynthesis rate at nitrogen content = 0. AMAXB is the increase of photosynthesis with increase of nitrogen percentage.
For parametrization, there are three different sources that have defined general values for broadleaved trees. These are Aber et al. 1995 (AMAXA: -46, AMAXB: 71.9, defined from investigations on Quercus rubra). Reich et al. 1995 (AMAXA 0.31*; AMAXB 5.45, defined from oak, maple and birch leaves). Kattge et al. 2009 (AMAXA: 5.73; AMAXB: 29.81 * 10.0 / SLAMIN, defined from a literature review with many different species).
The general practice now is to take the AMAXA from the newest and most elaborated dataset (Kattge et al. = 5.73) and adjust AMAXB according to the specific leaf area value at the top of the canopy. As long as there is no indication of SLAMIN, a standard of 18.1 is used (resulting from the assumption SLAMIN = 16.5).
References Photosyn().
Referenced by Photosyn(), and solve().

|
inline |
| lerr_t ldndc::PhysiologyPNET::solve | ( | ) |
@fixme The following two variables need to re-initialized after land use change. However, currently only arrays can be re-initialized in the treedyn module. Therefore, pnet can not be run for re-planting within the year after harvest (possible work-around: harvest at 31.12. of the year).
References AllocateMo(), AllocateYr(), N_uptake(), Phenology(), Photosyn(), ldndc::potential_wood_transpiration(), solve(), TransformPhysiology(), Update(), update_nitrogen_concentration(), and YearlyReset().
Referenced by solve().

|
private |
TransformPhysiology
References get_na_max_pnet(), ldndc::sap_wood_fraction(), SRATEMAX, and TransformPhysiology().
Referenced by AllocateMo(), solve(), and TransformPhysiology().


|
private |
Update
References Update().
Referenced by AllocateMo(), solve(), and Update().


|
private |
update_nitrogen_concentration
References update_nitrogen_concentration().
Referenced by solve(), and update_nitrogen_concentration().

|
private |
Resets:
References get_na_max_pnet(), and YearlyReset().
Referenced by solve(), and YearlyReset().
|
staticprivate |
parameter from frost hardiness function
|
staticprivate |
fraction of uncovered area that can be reoccuppied in one year
Referenced by AllocateYr().
|
staticprivate |
??
Referenced by N_uptake().
|
staticprivate |
??
Referenced by N_uptake().
|
staticprivate |
fraction of nitrogen from total deposition that is taken up by the canopy
|
staticprivate |
??
Referenced by TransformPhysiology().
|
staticprivate |
Optimum temperature where maximum N uptake occurs (Gessler et al. 1998)
Referenced by N_uptake().
|
staticprivate |
Minimum temperature where N uptake is possible (estimated from Gessler et al. 1998, corroborated by Alvarez-Uria and Koerner 2007)
Referenced by N_uptake().