 |
LandscapeDNDC 1.37.0
|
 |
LandscapeDNDC 1.37.0
|
Spatially explicit groundwater model. More...
Classes | |
| class | EcHy |
| Hydrology model EcHy. More... | |
| class | PhysiologyGrowthPSIM |
| Vegetation model GrowthPSIM. More... | |
| class | PhysiologyPlaMox |
| Vegetation model PlaMox. More... | |
| class | PhysiologyPNET |
| Vegetation model PNET. More... | |
| class | SoilChemistryMeTrX |
| Biogeochemical model MeTrx. More... | |
| class | WatercycleDNDC |
| Watercycle model WatercycleDNDC. More... | |
Enumerations | |
| enum | lmodule_flag_e { LMOD_FLAG_NONE = 0u , LMOD_FLAG_CONTROL = 1u << 0 , LMOD_FLAG_USER = 1u << 1 , LMOD_FLAG_SYSTEM = 1u << 2 , LMOD_FLAG_INITIALIZER = 1u << 3 , LMOD_FLAG_FINALIZER = 1u << 4 , LMOD_FLAG_MAINTENANCE = 1u << 5 , LMOD_FLAG_OUTPUT = 1u << 6 , LMOD_FLAG_TEST_MODULE = 1u << 7 , LMOD_FLAG_CNT_ , LMOD_FLAG_CNT = 1 + 1 + cbm::log2_t< LMOD_FLAG_CNT_-1 >::LOG2 } |
| enum | { LSUB_FLAG_NONE = 0u } |
| enum | |
| Lower bounds for number of foliage layers. | |
| enum | stomatalconductance_method_e |
| Stomatal conductance methods. | |
| enum | { CPOOL_VL , CPOOL_L , CPOOL_R , CPOOL_B , CPOOL_CNT } |
| Order of carbon pools in array passed to allocate_litter. More... | |
Functions | |
| lerr_t | assign_soillayer_porosity (double &_porosity_with_stones, size_t const &_nd_stratum, double const &_depth, double const &_stone_fraction, double const &_c_org_without_stones, double const &_bulk_density_without_stones, const ldndc::site::input_class_site_t *_site, const ldndc::soillayers::input_class_soillayers_t *_soillayers, const ldndc::soilparameters::input_class_soilparameters_t *_soilparameters, bool _info=false) |
| lerr_t | assign_soillayer_macropores (double &_macropores_with_stones, double &_wc_fc, double &_wc_sat, size_t const &_nd_stratum, double const &_depth, double const &_stone_fraction, double const &_porosity_with_stones, const ldndc::site::input_class_site_t *_site, const ldndc::soillayers::input_class_soillayers_t *_soillayers, const ldndc::soilparameters::input_class_soilparameters_t *_soilparameters, bool _info=false) |
| double | resistance_aero_abovecanopy (double const &, double const &) |
| returns boundary layer resistance above canopy | |
| double | resistance_aero_incanopy (double const &, double const &, double const &) |
| returns boundary layer resistance within canopy | |
| double | crown_shape_parameter (double _length, double lref, double ps) |
| Returns a crown shape parameter which is modified by canopy length. | |
| void | canopy_biomass_distribution (size_t const _fl_cnt_max, size_t const _fl_cnt, double const _ps, double const _h_bottom, double const _h_top, double const *_h_fl, double *_fFol_fl) |
| Foliage biomass distribution vertically across canopy. | |
| void | canopy_lai_distribution (double _sla_min, double _sla_max, double _height_max, double _height_min, double _mFol, double const *_fFol_fl, double const *_h_fl, double *_lai_fl, double *_sla_fl, size_t _foliage_layer_cnt, size_t _foliage_layer_cnt_max) |
| Leaf area distribution vertically across the whole canopy. | |
| void | estimate_single_layer_biomass (double const _ps, double const _h_bottom, double const _h_top, double _lw, double &_fract, double _h_cum_bottom, double _h_cum_top) |
| Vertical biomass estimation. | |
| void | fineroots_biomass_distribution (size_t _sl_cnt_litter, size_t _sl_cnt_soil, double _ps, double _rooting_depth, double *_f_frt_sl, double const *_h_sl) |
| double | get_optimum_sap_wood_biomass (MoBiLE_Plant *_p) |
| double | get_sapwood_foliage_ratio (MoBiLE_Plant *_p) |
| double | sapwood_foliage_area_ratio (double _height, double _qsf_p1, double _qsf_p2) |
| double | stomatal_conductance_ballberry_1987 (double, double, double, double) |
| Stomatal conductance according to Berry, Woodward and Ball 1987. | |
| double | stomatal_conductance_leuning_1990 (double, double, double, double, double) |
| Stomatal conductance according to Leuning 1990. | |
| double | stomatal_conductance_leuning_1995_a (double, double, double, double, double, double, double) |
| Stomatal conductance according to Leuning 1995. | |
| double | stomatal_conductance_eller_2020 (double, double, double, double, double, double, double, double, double, double, double) |
| Stomatal conductance according to Eller et al. 2020. | |
| double | potential_wood_transpiration (soillayers::input_class_soillayers_t const &, double _vpd, double _carbon_uptake, double _f_area, double _wuecmax, double _wuecmin, lvector_t< double > _h_sl, lvector_t< double > _wc, lvector_t< double > _wc_min, lvector_t< double > _wc_max) |
| Returns potential transpiration in [m] for trees. | |
| double | potential_crop_transpiration (double _co2, double _carbon_uptake, double _wuecmax) |
| Returns potential transpiration in [m] for crops and grass. | |
| double | potential_transpiration (size_t, double gsmin, double gsmax, double *_lai_fl, lvector_t< double > _vpd_fl, double *_relative_conductance_fl) |
| Returns potential transpiration in [m] for any species type. | |
| lerr_t | get_voc_emission_factors (ldndc::MoBiLE_Plant *, int, bool, double, int, double, double) |
| lerr_t LDNDC_API | allocate_litter (double _c, double _n, double _RCNRVL, double _RCNRL, double _RCNRR, double _C[], double &_n_diff) |
| calculates the amount of C which is added to the very labile, labile and recalcitrant decomposition pool | |
| double LDNDC_API | cation_exchange_capacity (double _clay, double _sand, double _som, double _ph, ecosystem_type_e _ecosystem) |
| Detailed description provided here . | |
| double | d_eff_air_a (double const &, double const &) |
| Returns effective diffusion coefficient depending on gas content. | |
| double | d_eff_air_b (double const &, double const &, double const &, double const &) |
| Returns effective diffusion coefficient depending on gas content and porosity. | |
| double | d_eff_air_millington_and_quirk_1961 (double const &, double const &) |
| Returns effective diffusion coefficient after [55]. | |
| lerr_t | update_litter_height (soillayers::input_class_soillayers_t const &, substate_watercycle_t &, substate_soilchemistry_t &) |
| Detailed description provided here . | |
| bool | forest_floor (size_t, soillayers::input_class_soillayers_t const &, const ldndc::site::input_class_site_t &) |
| First floor defined by: | |
| double LDNDC_API | sap_wood_fraction (double _lai_max, double _qsfa, double _height_min, double _height_max, double _depth) |
| sap_wood_fraction The calculation of the fraction of sapwood in coarse roots is based on the idealized volume fraction and assumes that the stem in the crown as well as the coarse roots are shaped as cones, and that core wood reaches a third of the cone | |
| lerr_t | wood_initialization (soillayers::input_class_soillayers_t const &, MoBiLE_Plant *, substate_soilchemistry_t &, ecosystem_type_e) |
| Initialization of aboveground and belowground wood debris, derived from existing vegetation. | |
| double | non_stomatal_water_limitation (double const &_var, double const &_var_ref, double const &_var_scale) |
| Non stomatal water limitation. | |
| double LDNDC_API | thornthwaite_heat_index (double, size_t, double, double) |
| Thornthwaite heat index. | |
| double LDNDC_API | thornthwaite (double, double, double, double, double) |
| Potential evapotranspiration after [65]. | |
Variables | |
| LDNDC_API lflags_t const | TMODE_DEFAULTS = TMODE_SUBDAILY|TMODE_PRE_DAILY|TMODE_POST_DAILY|TMODE_PRE_YEARLY|TMODE_POST_YEARLY |
Spatially explicit groundwater model.
data synthesizer base class
declare common types related to species
common types related to species (implementation)
declare species groups known to the simulation system
container class for one complete soil and humus parameters set
container class for site parameters
Declare common types related to site input.
declare common types related to setup input
common types related to setup input
declare common types related to remotesensing input
collection of various helper functions
class containing full set of van Genuchten parametrization
declare common types related to groundwater input
declare common types related to geometry input
common event related data types
(management) event factories
base class for (management) events
declare common types related to climate input
declare common types related to checkpoint input
declare common types related to air chemistry input
for now, also pulls in all kernel specific input declarations
| anonymous enum |
| lerr_t ldndc::allocate_litter | ( | double | _c, |
| double | _n, | ||
| double | _RCNRVL, | ||
| double | _RCNRL, | ||
| double | _RCNRR, | ||
| double | _C[], | ||
| double & | _n_diff ) |
calculates the amount of C which is added to the very labile, labile and recalcitrant decomposition pool
The needed information in this function are the amount of carbon in litter _c and the corresponding CN ratio (determined from nitrogen _n).
four cases: If the CN ratio is lower than the CN ratio of the very labile litter pool, all litter-C goes to the very labile decomposition pool.
If the CN ratio is between RCNRVL and the harmonic mean of RCNRVL and RCNRR, an increasing CN ratio decreases C_0 and increases C_1 and C_2.
If thc CN ratio is between the harmonic mean of RCNRVL and RCNRR and RCNRR, C_1 and C_2 decrease and only C_0 increases.
If CN ratio is higher than RCNRR, all Litter-C goes to C_2.
| _c | Carbon in litter flux |
| _n | Nitrogen in litter flux |
| _C | Organic carbon that is added to the
|
| _n_diff |
| lerr_t ldndc::assign_soillayer_macropores | ( | double & | _macropores_with_stones, |
| double & | _wc_fc, | ||
| double & | _wc_sat, | ||
| size_t const & | _nd_stratum, | ||
| double const & | _depth, | ||
| double const & | _stone_fraction, | ||
| double const & | _porosity_with_stones, | ||
| const ldndc::site::input_class_site_t * | _site, | ||
| const ldndc::soillayers::input_class_soillayers_t * | _soillayers, | ||
| const ldndc::soilparameters::input_class_soilparameters_t * | _soilparameters, | ||
| bool | _info = false ) |
Hydraulic parameters are checked for consistency against each other.
References forest_floor().
| lerr_t ldndc::assign_soillayer_porosity | ( | double & | _porosity_with_stones, |
| size_t const & | _nd_stratum, | ||
| double const & | _depth, | ||
| double const & | _stone_fraction, | ||
| double const & | _c_org_without_stones, | ||
| double const & | _bulk_density_without_stones, | ||
| const ldndc::site::input_class_site_t * | _site, | ||
| const ldndc::soillayers::input_class_soillayers_t * | _soillayers, | ||
| const ldndc::soilparameters::input_class_soilparameters_t * | _soilparameters, | ||
| bool | _info = false ) |
Porosity is estimated based on organic carbon fraction and bulk density for the soil including stones. Porosity is given by a value between 0 and 1. See: porosity()
| void ldndc::canopy_biomass_distribution | ( | size_t const | _fl_cnt_max, |
| size_t const | _fl_cnt, | ||
| double const | _ps, | ||
| double const | _h_bottom, | ||
| double const | _h_top, | ||
| double const * | _h_fl, | ||
| double * | _fFol_fl ) |
Foliage biomass distribution vertically across canopy.
| _fl_cnt_max | maximum number of foliage parameters |
| _fl_cnt | current actively used number of foliage parameters |
| _ps | |
| _h_bottom | |
| _h_top | |
| _h_fl | |
| _fFol_fl |
References estimate_single_layer_biomass().
| void ldndc::canopy_lai_distribution | ( | double | _sla_min, |
| double | _sla_max, | ||
| double | _height_max, | ||
| double | _height_min, | ||
| double | _mFol, | ||
| double const * | _fFol_fl, | ||
| double const * | _h_fl, | ||
| double * | _lai_fl, | ||
| double * | _sla_fl, | ||
| size_t | _foliage_layer_cnt, | ||
| size_t | _foliage_layer_cnt_max ) |
Leaf area distribution vertically across the whole canopy.
| _sla_min | specific leaf area at canopy top |
| _sla_max | specific leaf area at canopy bottom |
| _height_max | total plant height |
| _height_min | plant height at start of canopy |
| _mFol | foliage biomass |
| _fFol_fl | foliage biomass distribution |
| _h_fl | canopy discretization |
| _lai_fl | canopy layer leaf area index |
| _sla_fl | canopy layer specific leaf are |
| _foliage_layer_cnt | number of canopy layers |
| _foliage_layer_cnt_max | maximum number of canopy layers |
| double LDNDC_API ldndc::cation_exchange_capacity | ( | double | _clay, |
| double | _sand, | ||
| double | _som, | ||
| double | _ph, | ||
| ecosystem_type_e | _ecosystem ) |
Detailed description provided here .
| _clay | Clay content [%] |
| _sand | Sand content [%] |
| _som | Soil organic matter content [%] |
| _ph | pH value [-] |
| _ecosystem | Ecosystem type [-] |
Referenced by ldndc::SoilChemistryMeTrX::MeTrX_clay_nh4_equilibrium().
| double ldndc::crown_shape_parameter | ( | double | _length, |
| double | lref, | ||
| double | ps ) |
Returns a crown shape parameter which is modified by canopy length.
modification of crown shape by canopy length
| double ldndc::d_eff_air_a | ( | double const & | _exp, |
| double const & | _gas_content ) |
Returns effective diffusion coefficient depending on gas content.
| [in] | _exp | Exponent [-] |
| [in] | _gas_content | Gas content [m^3:m^-3] |
| double ldndc::d_eff_air_b | ( | double const & | _exp_g, |
| double const & | _exp_p, | ||
| double const & | _gas_content, | ||
| double const & | _porosity ) |
Returns effective diffusion coefficient depending on gas content and porosity.
| [in] | _exp_g | Exponent for gas content [-] |
| [in] | _exp_p | Exponent for porosity [-] |
| [in] | _gas_content | Gas content [m^3:m^-3] |
| [in] | _porosity | Porosity [m^3:m^-3] |
| double ldndc::d_eff_air_millington_and_quirk_1961 | ( | double const & | _gas_content, |
| double const & | _porosity ) |
Returns effective diffusion coefficient after [55].
| [in] | _gas_content | Gas content [m^3:m^-3] |
| [in] | _porosity | Porosity [m^3:m^-3] |
| void ldndc::estimate_single_layer_biomass | ( | double const | _ps, |
| double const | _h_bottom, | ||
| double const | _h_top, | ||
| double | _lw, | ||
| double & | _fract, | ||
| double | _h_cum_bottom, | ||
| double | _h_cum_top ) |
Vertical biomass estimation.
| _ps | form parameter |
| _h_bottom | height of start of distributed biomass |
| _h_top | height of end of distributed biomass |
| _lw | |
| _fract | |
| _h_cum_bottom | |
| _h_cum_top |
The standard vertical distribution of biomass is done with the same assumptions for canopy (veglibs_canopy) and rooting space (Root distribution). Only for root distribution there are several more options available.
Referenced by canopy_biomass_distribution(), and fineroots_biomass_distribution().

| void ldndc::fineroots_biomass_distribution | ( | size_t | _sl_cnt_litter, |
| size_t | _sl_cnt_soil, | ||
| double | _ps, | ||
| double | _rooting_depth, | ||
| double * | _f_frt_sl, | ||
| double const * | _h_sl ) |
Root distribution and root properties are generally fixed throughout the simulation period. Only, if the initial condition of planting doesn't allow the soil profile to be fully rooted (e.g. plants are too small), rooting depth increases linearly with \( GZRTZ \) until the maximum root depth (defined by the soil profile) is reached. This is different in Plamox, where plant roots start growing down with planting and should not grow as deep as the soil profile depth, but stop at the species parameter, the maximum rooting depth \( ZRTMC \).
The distribution of fine root biomass \( m(z) \) within the rooting space depends on the rooting depth and is calculated with the same empirical equation as the foliage biomass (veglibs_canopy). The main difference is that the parameter 'p' is generally below 1 now to get a concave distribution ([28]).
\[ m(z) = M \cdot \frac{r_{ih} f(z)}{\int r_{ih} f(z)} \\ r_{ih} = \frac{RD - z}{RD} \\ f(z) = p^{100 \frac{z}{RD^2}} \]
\( RD \): root depth (m)
\( z \): distribution depth for which the percentage roots should be defined (between 0-1, from upper soil boundary to rooting depth)
\( p \): shape parameter
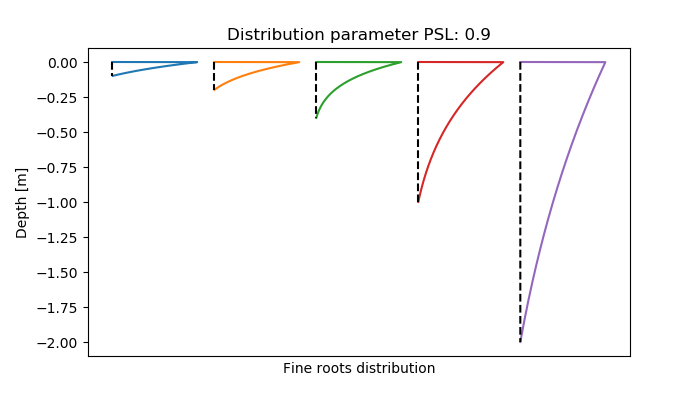
scaling factor for root distribution (basically to enable root and canopy profile calculations with the same function)
fraction of fine roots in the litter layer
fraction of fine roots in the mineral soil
References estimate_single_layer_biomass().

| bool ldndc::forest_floor | ( | size_t | _sl, |
| soillayers::input_class_soillayers_t const & | _soillayers, | ||
| const ldndc::site::input_class_site_t & | ) |
First floor defined by:
Referenced by assign_soillayer_macropores().

| double ldndc::get_optimum_sap_wood_biomass | ( | MoBiLE_Plant * | _p | ) |
| _p | Plant |
Referenced by get_sapwood_foliage_ratio().

| double ldndc::get_sapwood_foliage_ratio | ( | MoBiLE_Plant * | _p | ) |
| _p | Plant |
References get_optimum_sap_wood_biomass().

| lerr_t ldndc::get_voc_emission_factors | ( | ldndc::MoBiLE_Plant * | _vt, |
| int | _fl, | ||
| bool | _calc_synthase_activity, | ||
| double | _latitude, | ||
| int | _yearday, | ||
| double | _foliage_temperature, | ||
| double | _foliage_radiation ) |
Daily calculation of voc (isoprene and monoterpenes) synthase activity NOTE that fCO2 depends on outside CO2, internal CO2 would probably be better (explaining relatively high emissions when stomata closed)
| double ldndc::non_stomatal_water_limitation | ( | double const & | _var, |
| double const & | _var_ref, | ||
| double const & | _var_scale ) |
Non stomatal water limitation.
| _var | ... |
| _var_ref | ... |
| _var_scale | ... |
Referenced by ldndc::PhysiologyGrowthPSIM::PSIM_PhotosynthesisRates().

| double ldndc::sap_wood_fraction | ( | double | _lai_max, |
| double | _qsfa, | ||
| double | _height_min, | ||
| double | _height_max, | ||
| double | _depth ) |
sap_wood_fraction The calculation of the fraction of sapwood in coarse roots is based on the idealized volume fraction and assumes that the stem in the crown as well as the coarse roots are shaped as cones, and that core wood reaches a third of the cone
Calculation is based on the sapwood area demand for foliage. Sapwood volume is assumed to equal a cone within the canopy and across the rooted soil, and a column between the ground and canopy start.
Referenced by ldndc::PhysiologyPNET::TransformPhysiology().

| double ldndc::sapwood_foliage_area_ratio | ( | double | _height, |
| double | _qsf_p1, | ||
| double | _qsf_p2 ) |
| _height | Tree height |
| _qsf_p1 | species parameter |
| _qsf_p2 | species parameter |
| lerr_t ldndc::update_litter_height | ( | soillayers::input_class_soillayers_t const & | _soillayers, |
| substate_watercycle_t & | _wc, | ||
| substate_soilchemistry_t & | _sc ) |
Detailed description provided here .
| _sl | Soil layers class |
| _wc | Watercycle state |
| _sc | Soilchemistry state |
| lerr_t ldndc::wood_initialization | ( | soillayers::input_class_soillayers_t const & | _sl, |
| MoBiLE_Plant * | _p, | ||
| substate_soilchemistry_t & | _sc, | ||
| ecosystem_type_e | _ecosystemtype ) |
Initialization of aboveground and belowground wood debris, derived from existing vegetation.
Referenced by ldndc::SoilChemistryMeTrX::MeTrX_update().

|
extern |